We have used automated systems with oscillating microliter droplets to screen and optimize thermally driven, electrochemical, and photochemical reactions. Oscillating the individual droplets promote mixing regardless of reaction time but increases challenges in controlling evaporation. Each droplet behaves as a batch reactor, but the advantages over batch reactions in a well plate or small vials are (1) individual control of reaction conditions, including photon flux for photochemistry, (2) reproducible mixing, (3) access to elevated temperature and pressure, and (4) ability to handle biphasic reactions. Moreover, droplets enable multiobjective optimization of reaction conditions over continuous variables (e.g., temperature, time, and concentrations) and categorical parameters (e.g., solvents, ligands, and catalysts). Current efforts extend the approach to multiple independent parallel droplet units, increasing throughput since each droplet can operate in different conditions from its neighbors. The ability to run reactions in parallel, independently controlled droplets enhances the platform’s performance in kinetic studies and optimization of reaction conditions.
A critical aspect of feedback optimization of reactions is automated, reliable online analytics, typically by HPLC/MS. Overlapping peaks or inaccurate integration routines lead to wrong analytical results returned to the experimental design algorithm and interruption of the automated optimization cycle. We have developed an open-source Python project, MOCCA, to analyze the raw data of HPLC–DAD (photodiode array detector). It provides several analysis features, including an automated peak deconvolution routine of known signals, even if overlapped with signals of unexpected impurities or side products. As one of the application examples of MOCCA, we demonstrated closed-loop optimization of the alkylation of 2-pyridone. The Bayesian optimizer (EDBO) proposed new experimental parameter values and fed them into a LabVIEW program controlling our microfluidic droplet platform. After the reaction, the droplet was diluted with acetonitrile and moved to an internal injection valve to inject a sample directly into the HPLC system. After each run, the HPLC system automatically exported HPLC–DAD raw data for MOCCA data analysis, which delivered the reaction outcome to the optimizer, closing the cycle.
Selected publications:
- L.M. Baumgartner, J.M. Dennis, N.A. White, S.L. Buchwald, and K.F. Jensen. Use of a droplet platform to optimize Pd-catalyzed C-N coupling reactions promoted by organic bases. Org. Process Res. & Dev., 23 (8) 1594-1601 (2019).
- H.-W. Hsieh, C. W. Coley, L. Baumgartner, K. F. Jensen, R. Robinson, Photoredox iridium-nickel dual catalyzed decarboxylative arylation cross-coupling: from batch to continuous flow via self-optimizing segmented flow reactor. Org. Process Res. Dev, 22 (4), 542–550 (2018).
- C. P. Haas, M. Lübbesmeyer, E. Jin, M.A McDonald, B. Koscher, N. Guimond, L. Di Rocco, H. Kayser, S. Leweke, S. Niedenführ, R. Nicholls, E. Greeves, D. Barber, J. Hillenbrand, G. Volpin, and K.F. Jensen, Open-Source Chromatographic Data Analysis for Reaction Optimization and Screening, accepted ACS Cent. Sci.
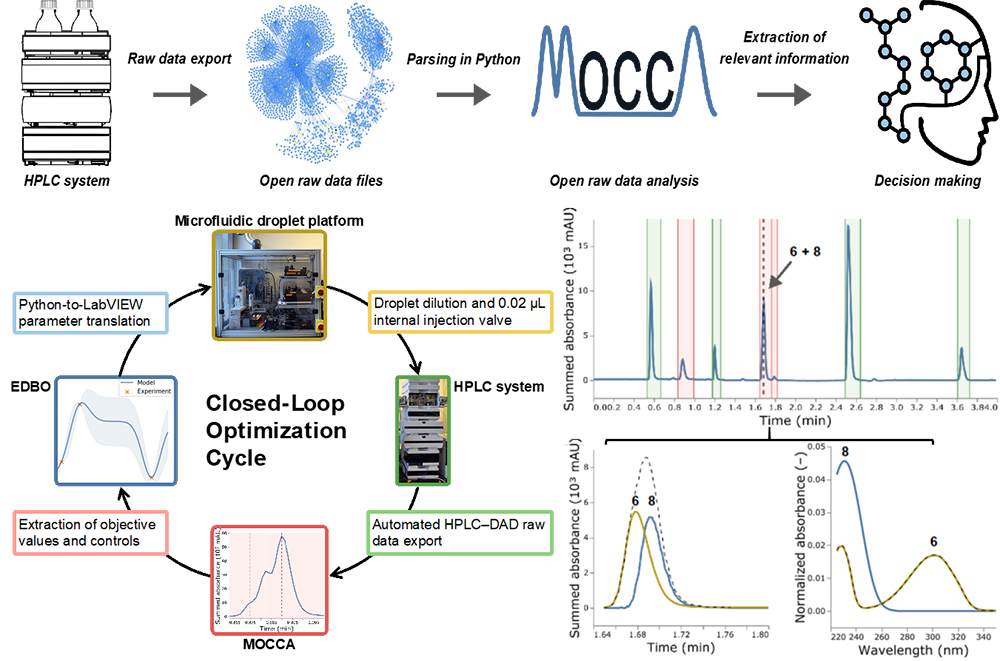
Top: HPLC analysis workflow. HPLC–DAD raw data are exported in non-proprietary and open data formats. After parsing in Python, HPLC–DAD datasets are analyzed in context to each other by MOCCA, and structured datasets are generated for data-based decision-making. Bottom left: Closed-loop optimization cycle. Blue: Experimental Design via Bayesian Optimization (EDBO) Python package and translation of the suggested parameters to a LabVIEW experimental protocol. Yellow: Experimental execution by a microfluidic reactor platform employing an oscillatory droplet reactor design. 0.02 μL HPLC samples are taken out of the droplet after dilution with acetonitrile. Green: HPLC system with a photodiode array detector (DAD) and an automated HPLC–DAD raw data export routine. Red: Data analysis by the MOCCA tool and a project-specific script for the extraction of objective values and process control values. Right middle: Chromatogram of the reaction under optimal conditions with an impure product peak (~1.7 min). Right bottom: Modelled retention profiles (dashed line: impure peak) and UV-Vis spectra (dashed line: reference UV-Vis spectrum of 6) of the pure product 6 (yellow) and the unexpected impurity 8 (blue).
C. P. Haas, et al., ACS Cent. Sci.

