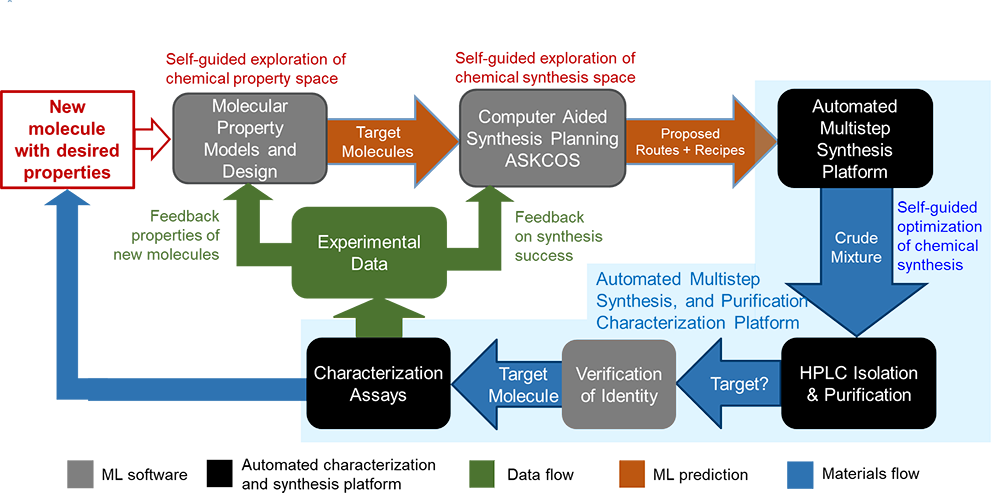
Based on our experiences with automated reaction systems and computer-aided synthesis planning, we have spent the past 3+ years developing an automated platform integrating generative machine learning (ML) models and property prediction ML models from MIT colleagues (Prof. Barzilay, Green, Gomez-Bombarelli, and Jaakkola) with ASKCOS, our synthesis models, Mongo database, master control system, and Bayesian optimization to enable autonomous molecular discovery. Dyes serve as demonstration cases because of their rich chemical diversity and wide-ranging applications, including molecular dyes for LED displays, biological labels, paints, and solar cell sensitizers and absorbers. Furthermore, they have the advantage that the desired properties (e.g., color, toxicity, and stability) are readily measured. Notably, the chemistry of dyes is diverse and sufficiently complex to allow the development of general ML methods and experimental techniques.
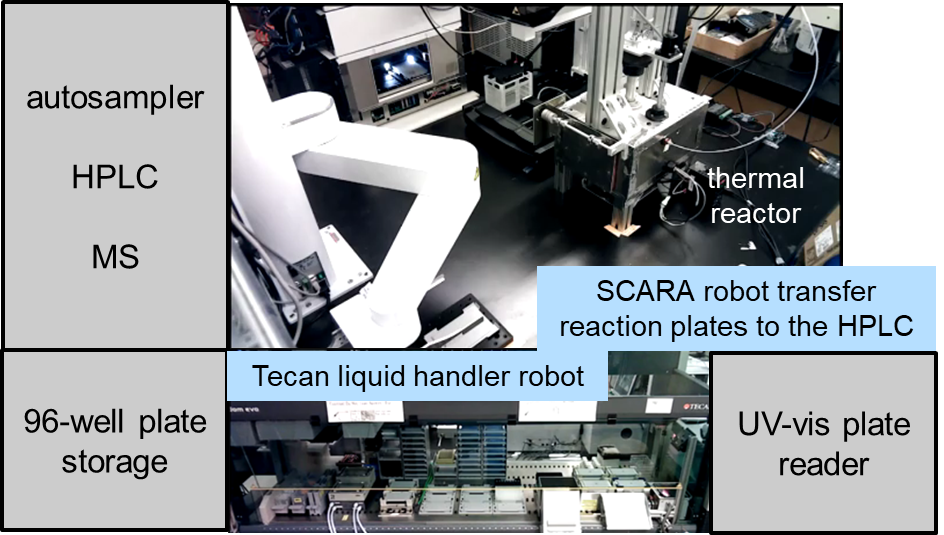
We have demonstrated the ability (1) to use generative models to propose new molecules not previously reported and having desired properties, (2) to automatically synthesize, purify, and characterize these molecules, and then (3a) to update the predictive ML models for a subsequent cycle of explorations, or (3b) to perform automated synthesis of a final set of molecules exploiting the refined ML models. Because the discovery process can involve solids, we perform multistep synthesis in batch, using a 96-well format with a TECAN liquid handler robot and customized reaction and separation platforms. ASKCOS proposes synthesis pathways; rule-based algorithms and the use of optimization mitigate shortfalls in context recommendations. Automated HLPC and MS purify and identify products whose logP, UV-vis, and oxidative degradation are subsequently automatically characterized. Additional current projects in the platforms include photochemistry, biological assays, and elucidating chemical mechanisms of catalytic reactions.
Publication:
- B.A. Koscher, R.B. Canty, M.A. McDonald, K.P. Greenman, C.J. McGill, C.L. Bilodeau, W.Jin, H. Wu, F.H. Vermeire, B. Jin, T. Hart, T. Kulesza, S.-C. Li, T.S. Jaakkola, R. Barzilay, R. Gómez-Bombarelli, W.H. Green, K.F. Jensen, Autonomous, multiproperty-driven molecular discovery: From predictions to measurements and back, Science 382, eadi1407 (2023). DOI: 10.1126/science.adi1407